- Blog
- About
- Contact
- Photograv troubleshooting
- Pirates of the caribbean the curse of the black pearl free
- Insert disk in drive d mimio studio
- Deleting base pairs in bioedit
- Winclone 5 download free
- How to build a hidden markov model matlab
- Airline guitar 1961
- Eternal poison faq
- Parallels vs vmware for visual studio
- Xojo keygen
- Delay analysis in primavera p6 pdf
- Daz studio genesis anime
- Rab ne bana di jodi movie english subtitles
- Jennifer lopez on the floor live on wetten dass
- Softraid for windows 10
- Pcsx2 1-4-0 bios reddit
- Corel draw x6 tutorials
- Silsilah keluarga korea
- Would you rather questions funny
- Delft university data orcaflex matlab
- Git create branch and push to server
- Techstream program key

North Carolina State University, Department of Microbiology. Hit save control-shift-s and repeat for each group of sequences. You may have to scroll down the program list to find it. Save the reverse complement as a text file under a different name. Then I run a NJ analysis to see what is going on with the dataset. This will allow you to see any base pairs that are different in the clean forwards.
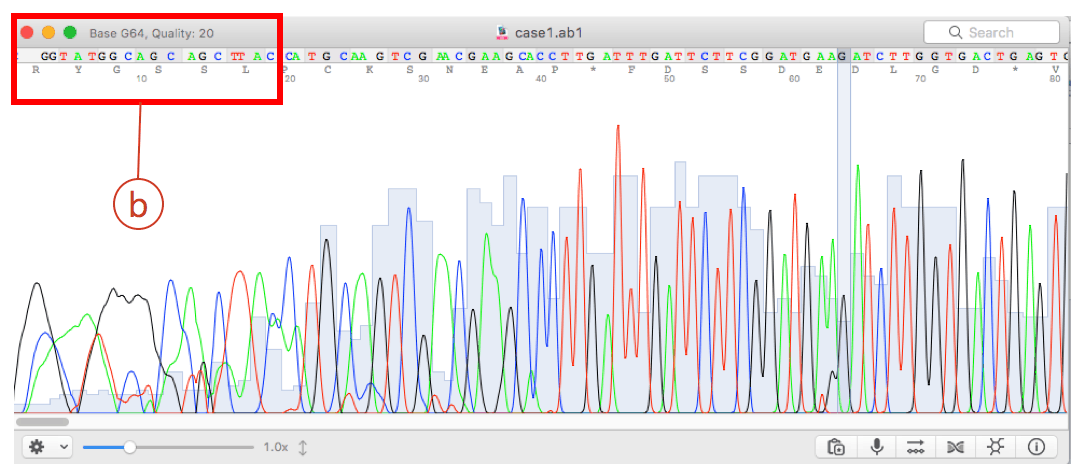
Guide to editing sequences with Chromas and BioEdit Select from the next next residue to the end. Once you set your preferences on one machine you can copy the bioedit. You should be able to clearly see the peaks of the trace. BioEdit Tutorials – Practical Bioinformatics Note that this works best with coding sequences without indels as every sequence is an identical length, it is all a bit trickier with different length sequences. I use this feature on nearly every dataset I create. BioEdit lets you modify just about anything that it does relative to menus and keyboard short cuts as well as the default settings for displaying data. Note how many replacements it does, this is the number of samples. Change the view type on the lower toolbar 3rd of the alignment windowselect the third colored button from the left says Shade identities and similarities when you hold the mouse over it. I always keep the BioEdit file with all forwards, reverses and consensus sequences so that if I double check stuff later it is easier to find the relevant chromatograms I can tell what sequence is from where by the sequence name. Move cursor between the residue and the previous residue. Much editing in BioEdit requires extensive repetitive actions, so using the menus will rather slow. In that case Nioedit try and get them close, but each individual one many require adjustment. At the end of this phase you have done two data checks, one when you edited your original chromatogram, second when you checked any unique base pair changes.Ĭlick on the edited forward sequence file to open it.

In BioEdit, clean up all the ends and get things to the base pairs you want to analyze. Then I undo the cut, select all the sequences Edit, Select All Sequences, control-shift-acopy them control-a–note that copy and pasting sequences is different to any other copy and paste action. The sequence present in the original file is the sequence of the newly synthesized strand.

My sequence names look like this, PU Select all residues of all the sequences. Select all the reverse sequences and cut them. If the vector sequence given is the opposite strand to the forward sequence, then there should be a region of almost exact homology with the end of the reverse complement. Now when you double click on a chromatogram it will open in Chromas. At that point I finish my consensus sequence. Select both files with the mouse by dragging it over the file names at the left. Delete and copy the data of highlighted sequence. Each line in the trace is colour-coded to match the colour that one of the 4 bases is displayed in. Save the file as text only and make sure it has the correct file extension. Select them all control-acopy to clipboard control-cgo back to BioEdit, to paste these names over the existing ones. Save the edited file to the desktop, or preferably, your own disk or network account. Select to the end including the current residue. There are 4 disks containing sequence files. Note that sequences after bases become increasingly unreliable, and are not worth spending much time on. This highlights any columns that have different bases. Now your BioEdit file has all the forwards and reverses, with the. The reason why I paste them to a new file first is that importing from the clipboard File, Import from Clipboard will place them at the bottom of your file, which is usually not where I want them be. Since this may interfere with analysis of the sequence, these will have to be edited out. These should show an almost exact match to the forward titorial reverse sequence. These are my preferences, you can use these or change them whatever you prefer.
Deleting base pairs in bioedit full#
See sequence analysis references for full map. MEGA also has an alignment editor, but I’ve not really used it very much. BioEdit can also edit chromatograms, but I find Chromas to be nicer. BioEdit is a mouse-driven, easy-to-use sequence alignment editor and sequence analysis program designed and written by a graduate student. This is likely to be the final release of BioEdit.
